SPC vs Wilcoxon
Delivering trusted, evidence-based answers from data—leaders can act on with confidence, using a time-tested method.
Understanding variation is one of the most underappreciated skills in data analysis.
Most analysts focus on models, tests, and dashboards.
Yet the real power lies in learning how a process behaves over time.
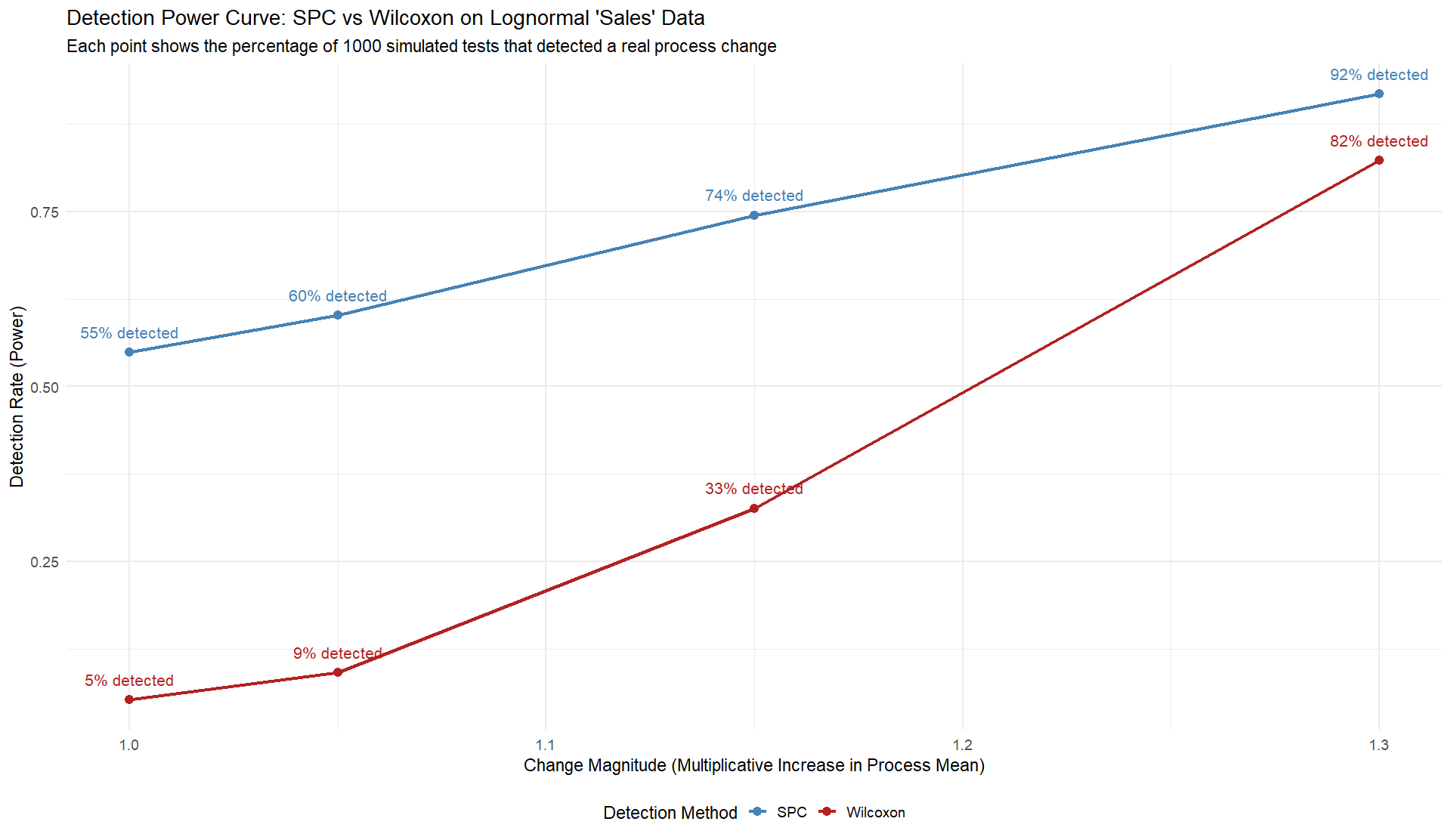
The comparison between the two detection methods—one focused on statistical testing and the other grounded in understanding variation—revealed something important: many analytical tools can signal change, but few can tell us whether that change truly means something.
In the simulation, code below, both methods started from the same data.
The difference wasn’t in the numbers but in how those numbers were interpreted.
The traditional statistical test identified differences between two samples, flagging significance whenever the distributions diverged beyond a threshold.
It’s precise, even elegant, but often blind to the process that produced the data.
By contrast, the method based on variation looked at the entire timeline.
It didn’t just ask whether two moments were different; it asked whether the process itself had shifted in a meaningful way.
That distinction matters because most of what we see in real-world data isn’t signal.
It’s noise.
Sales fluctuate.
Website visits spike and fall.
Production output drifts slightly day by day.
Without understanding variation, every small change can appear urgent, every dip a crisis, every increase a victory.
The simulation showed that when we treat all movement as meaningful, we make poor decisions most of the time.
It also showed that learning to recognize stable versus unstable behavior changes the way we act. Instead of reacting to noise, we begin to respond to signals.
Many businesses rely on statistical significance testing to guide their decisions.
It’s tempting because it provides clear answers: something changed or it didn’t.
But data is rarely that simple.
Statistical tests compare two sets of data as if they were isolated events.
They’re useful for experiments, not for ongoing processes.
In an operational context, the question isn’t whether two samples differ but whether a system has truly changed.
When we measure over time, we start to see the story behind the numbers.
Patterns emerge, cycles reveal themselves, and causes become clearer.
This understanding doesn’t come from more sophisticated tools; it comes from better thinking.
Recognizing variation means asking better questions of the data.
Is the change I’m seeing part of a pattern or an exception?
Is the variation predictable or special?
Can I trust this signal enough to act on it?
These questions shift analysis from a reactive to a reflective practice.
The results of the simulation highlight the practical consequences of this mindset.
One method detected change about three-quarters of the time when it truly existed.
The other detected it roughly half as often.
The difference wasn’t computational power; it was conceptual clarity.
One approach saw variation as a problem to eliminate, the other as a message to interpret.
Understanding variation allows us to make sense of uncertainty.
It helps prevent false alarms and missed opportunities.
It keeps us grounded in what the data can and cannot tell us.
This approach requires patience, discipline, and humility—the willingness to wait for patterns to reveal themselves before jumping to conclusions.
In the end, data analysis is about understanding behavior.
Variation provides the context that makes numbers meaningful.
When we learn to see it clearly, our analyses become less about testing and more about learning.
The lesson from this comparison is not which method is superior but how much more powerful analysis becomes when guided by an understanding of variation rather than a fixation on difference.
Try the R below to run this test in RStudio.
# ============================================================
# SPC vs Wilcoxon on lognormal "sales" + visualizations
# ============================================================
set.seed(999)
# Parameters
n_pre <- 30
n_post <- 30
sigma_logn <- 0.35
mu_base <- log(100) - 0.5 * sigma_logn^2
alpha <- 0.05
n_sims <- 1000
multipliers <- c(1.00, 1.05, 1.15, 1.30)
# Data generator
r_logn_sales <- function(n, mu_log=mu_base, sigma_log=sigma_logn, mult=1){
rlnorm(n, meanlog = mu_log + log(mult), sdlog = sigma_log)
}
# Phase I limits on log scale; return raw-scale bands too (for plotting)
phase1_limits <- function(x_pre) {
z <- log(x_pre)
mr <- abs(diff(z))
MRb <- mean(mr, na.rm = TRUE)
mu <- mean(z, na.rm = TRUE)
UCLz <- mu + 2.66 * MRb
LCLz <- mu - 2.66 * MRb
list(mean_log = mu, MRbar_log = MRb,
UCL_log = UCLz, LCL_log = LCLz,
mean_raw = exp(mu), UCL_raw = exp(UCLz), LCL_raw = exp(LCLz))
}
# SPC detection (three rules) using log-sized limits
spc_detect <- function(x_pre, x_post, run_len_rule2 = 8, outer_band_frac = 0.75) {
lim <- phase1_limits(x_pre)
z_post <- log(x_post)
mu <- lim$mean_log; UCL <- lim$UCL_log; LCL <- lim$LCL_log
# Rule 1
rule1 <- any(z_post > UCL | z_post < LCL, na.rm = TRUE)
# Rule 2
side_post <- ifelse(z_post > mu, 1L, ifelse(z_post < mu, -1L, 0L))
side_post[side_post == 0L] <- NA_integer_
r <- rle(na.omit(side_post))
rule2 <- if (length(r$lengths)) any(r$lengths >= run_len_rule2) else FALSE
# Rule 3
upper_outer <- mu + outer_band_frac * (UCL - mu)
lower_outer <- mu - outer_band_frac * (mu - LCL)
is_up <- z_post >= upper_outer
is_lo <- z_post <= lower_outer
rule3 <- FALSE
n <- length(z_post)
if (n >= 4) {
for (i in seq_len(n - 3)) {
if (sum(is_up[i:(i+3)], na.rm=TRUE) >= 3 ||
sum(is_lo[i:(i+3)], na.rm=TRUE) >= 3) { rule3 <- TRUE; break }
}
}
list(any_special = (rule1 || rule2 || rule3),
rule1 = rule1, rule2 = rule2, rule3 = rule3, limits = lim,
thresholds = list(upper_outer = exp(upper_outer), lower_outer = exp(lower_outer)))
}
# 1) Monte Carlo runner that returns detailed rows (for plotting)
# One Monte Carlo at a given change multiplier; returns row per sim
mc_rows_spc_vs_wilcox <- function(n_sims = 1000, change_mult = 1.00,
run_len_rule2 = 8, alpha = 0.05) {
out <- replicate(n_sims, {
x_pre <- r_logn_sales(n_pre, mult = 1.0)
x_post <- r_logn_sales(n_post, mult = change_mult)
# Wilcoxon rank-sum (non-parametric)
p_wilcox <- wilcox.test(x_post, x_pre, exact = FALSE)$p.value
wilcox_sig <- as.integer(p_wilcox < alpha)
# SPC
spc <- spc_detect(x_pre, x_post, run_len_rule2 = run_len_rule2)
spc_sig <- as.integer(spc$any_special)
c(spc = spc_sig, wilcox = wilcox_sig, p_wilcox = p_wilcox)
})
as.data.frame(t(out))
}
# Run across multipliers; bind for plotting
run_grid <- function(mult_vec = multipliers, n_sims = 1000, alpha = 0.05) {
dfs <- lapply(mult_vec, function(m) {
df <- mc_rows_spc_vs_wilcox(n_sims = n_sims, change_mult = m, alpha = alpha)
df$mult <- m
df
})
do.call(rbind, dfs)
}
res <- run_grid(multipliers, n_sims = n_sims, alpha = alpha)
# 2) Summaries you’ll plot
# Power (detection rate) by method and multiplier
power_summary <- function(res_df){
aggregate(. ~ mult, data = transform(res_df,
spc_det = as.numeric(spc==1),
wilcox_det = as.numeric(wilcox==1)),
FUN = mean)[, c("mult","spc_det","wilcox_det")]
}
pow <- power_summary(res)
# Medium-change confusion matrix (e.g., 1.15)
subset_medium <- subset(res, mult == 1.15)
cm <- table(Wilcoxon = factor(subset_medium$wilcox, levels=c(0,1), labels=c("No","Yes")),
SPC = factor(subset_medium$spc, levels=c(0,1), labels=c("No","Yes")))
cm_df <- as.data.frame(cm)
colnames(cm_df) <- c("Wilcoxon","SPC","Count")
cm_df$Percent <- cm_df$Count / sum(cm_df$Count)
# Null p-value calibration
subset_null <- subset(res, mult == 1.00)
#3) Visuals (ggplot2)
if (!requireNamespace("ggplot2", quietly = TRUE)) {
stop("Please install.packages('ggplot2') to produce the visuals.")
}
library(ggplot2)
library(tidyr)
# Your data
df_power <- data.frame(
change_mult = c(1.00, 1.05, 1.15, 1.30),
SPC = c(0.549, 0.602, 0.745, 0.918),
Wilcoxon = c(0.052, 0.091, 0.326, 0.823)
)
# Reshape for plotting
df_long <- pivot_longer(df_power, cols = c("SPC", "Wilcoxon"), names_to = "Method", values_to = "DetectionRate")
# Create annotation text for each point
df_long$label <- paste0(round(df_long$DetectionRate * 100), "% detected")
# Plot with annotations
ggplot(df_long, aes(x = change_mult, y = DetectionRate, color = Method)) +
geom_line(size = 1.2) +
geom_point(size = 3) +
geom_text(
aes(label = label),
vjust = -1.2,
size = 4.5,
show.legend = FALSE
) +
scale_color_manual(values = c("SPC" = "steelblue", "Wilcoxon" = "firebrick")) +
labs(
title = "Detection Power Curve: SPC vs Wilcoxon on Lognormal 'Sales' Data",
subtitle = "Each point shows the percentage of 1000 simulated tests that detected a real process change",
x = "Change Magnitude (Multiplicative Increase in Process Mean)",
y = "Detection Rate (Power)",
color = "Detection Method"
) +
theme_minimal(base_size = 14) +
theme(legend.position = "bottom")
Consultation